2D parameter estimation using MCMC#
This example will cover:
Use MCMC to infer kernel paramaters
Finding sample with highest log-prob from the mcmc chain
Visualising results of sampling
Making predictions
[1]:
from gptide import cov
from gptide import GPtideScipy
import numpy as np
import matplotlib.pyplot as plt
import corner
import arviz as az
from scipy import stats
from gptide import stats as gpstats
Generate some data#
[2]:
####
# These are our kernel input parameters
np.random.seed(1)
noise = 0.5
η = 10
ℓ_x = 900
ℓ_y = 1800
dx = 200.
dy = 400.
def kernel_2d(x, xpr, params):
"""
2D kernel
Inputs:
x: matrices input points [N,3]
xpr: matrices output points [M,3]
params: tuple length 3
eta: standard deviation
lx: x length scale
ly: y length scale
"""
eta, lx, ly = params
# Build the covariance matrix
C = cov.matern32(x[:,1,None], xpr.T[:,1,None].T, ly)
C *= cov.matern32(x[:,0,None], xpr.T[:,0,None].T, lx)
C *= eta**2
return C
covfunc = kernel_2d
###
# Domain size parameters
N = 10
covparams = (η, ℓ_x, ℓ_y)
# Input data points
xd = np.arange(0,dx*N,dx)[:,None]-dx/2
yd = np.arange(0,dy*N,dy)[:,None]-dy/2
# Make a grid
Xg, Yg = np.meshgrid(xd, yd)
# Vectorise grid and stack
Xv = Xg.ravel()
Yv = Yg.ravel()
X = np.hstack([Xv[:,None], Yv[:,None]])
GP = GPtideScipy(X, X.copy(), noise, covfunc, covparams)
# Use the .prior() method to obtain some samples
zd = GP.prior(samples=1)
zg = zd.reshape(Xg.shape)
[3]:
plt.figure()
plt.scatter(Xg, Yg, c=zg)
plt.ylabel('y')
plt.xlabel('x')
plt.title('Noisy Matern 3/2')
plt.colorbar(label='some data')
plt.gca().set_aspect('equal')
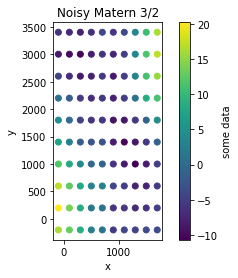
Inference#
We now use the gptide.mcmc function do the parameter estimation. This uses the emcee.EnsembleSampler class.
[4]:
from gptide import mcmc
n = len(xd)
covparams
[4]:
(10, 900, 1800)
[6]:
# Initial guess of the noise and covariance parameters (these can matter)
noise_prior = gpstats.truncnorm(0.4, 0.25, 1e-15, 1) # noise - true value 0.5
covparams_priors = [gpstats.truncnorm(8, 3, 2, 14), # eta - true value 10
gpstats.truncnorm(600, 200, 1e-15, 1e4), # ℓ_x - true value 900
gpstats.truncnorm(1400, 250, 1e-15, 1e4) # ℓ_y - true value 1800
]
samples, log_prob, priors_out, sampler = mcmc.mcmc( X,
zd,
covfunc,
covparams_priors,
noise_prior,
nwarmup=30,
niter=100,
verbose=False)
Running burn-in...
100%|██████████████████████████████████████████████████████████████████████████████████| 30/30 [00:16<00:00, 1.81it/s]
Running production...
100%|████████████████████████████████████████████████████████████████████████████████| 100/100 [00:56<00:00, 1.77it/s]
Find sample with highest log prob#
[7]:
i = np.argmax(log_prob)
MAP = samples[i, :]
print('Noise (true): {:3.2f}, Noise (mcmc): {:3.2f}'.format(noise, MAP[0]))
print('η (true): {:3.2f}, η (mcmc): {:3.2f}'.format(covparams[0], MAP[1]))
print('ℓ_x (true): {:3.2f}, ℓ_x (mcmc): {:3.2f}'.format(covparams[1], MAP[2]))
print('ℓ_y (true): {:3.2f}, ℓ_y (mcmc): {:3.2f}'.format(covparams[2], MAP[3]))
Noise (true): 0.50, Noise (mcmc): 0.29
η (true): 10.00, η (mcmc): 8.34
ℓ_x (true): 900.00, ℓ_x (mcmc): 705.59
ℓ_y (true): 1800.00, ℓ_y (mcmc): 1692.21
[8]:
i = np.argmax(log_prob)
MAP = samples[i, :]
print('Noise (true): {:3.2f}, Noise (mcmc): {:3.2f}'.format(noise, MAP[0]))
print('η (true): {:3.2f}, η (mcmc): {:3.2f}'.format(covparams[0], MAP[1]))
print('ℓ_x (true): {:3.2f}, ℓ_x (mcmc): {:3.2f}'.format(covparams[1], MAP[2]))
print('ℓ_y (true): {:3.2f}, ℓ_y (mcmc): {:3.2f}'.format(covparams[2], MAP[3]))
Noise (true): 0.50, Noise (mcmc): 0.29
η (true): 10.00, η (mcmc): 8.34
ℓ_x (true): 900.00, ℓ_x (mcmc): 705.59
ℓ_y (true): 1800.00, ℓ_y (mcmc): 1692.21
Posterior density plot#
[9]:
labels = ['σ','η','ℓ_x', 'ℓ_y']
def convert_to_az(d, labels):
output = {}
for ii, ll in enumerate(labels):
output.update({ll:d[:,ii]})
return az.convert_to_dataset(output)
priors_out_az = convert_to_az(priors_out, labels)
samples_az = convert_to_az(samples, labels)
axs = az.plot_density( [samples_az[labels],
priors_out_az[labels]],
shade=0.1,
grid=(1, 5),
textsize=12,
figsize=(12,3),
data_labels=('posterior','prior'),
hdi_prob=0.995)
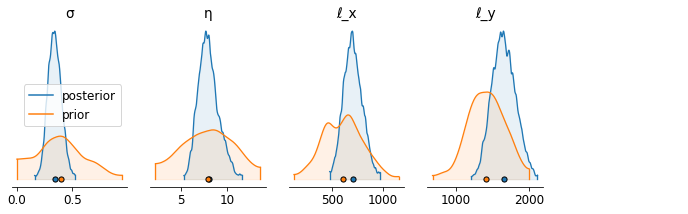
Posterior corner plot#
[10]:
fig = corner.corner(samples,
show_titles=True,
labels=labels,
plot_datapoints=True,
quantiles=[0.16, 0.5, 0.84])
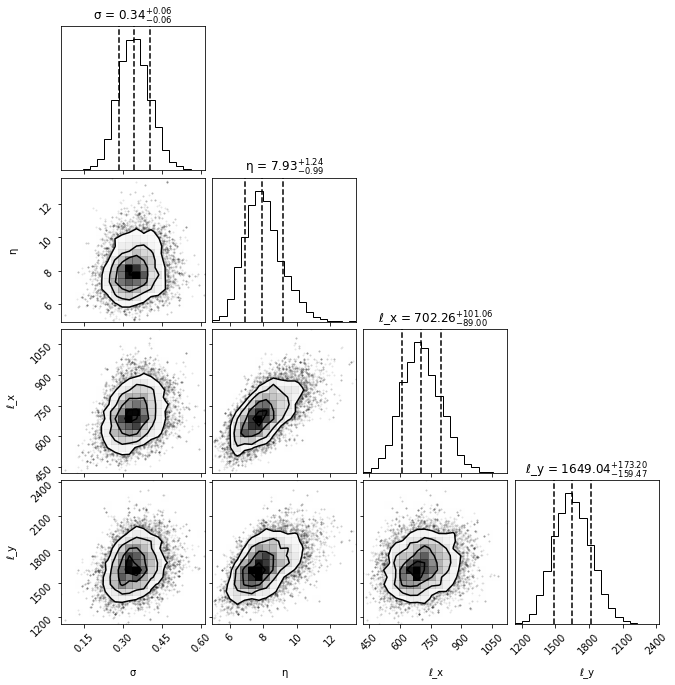
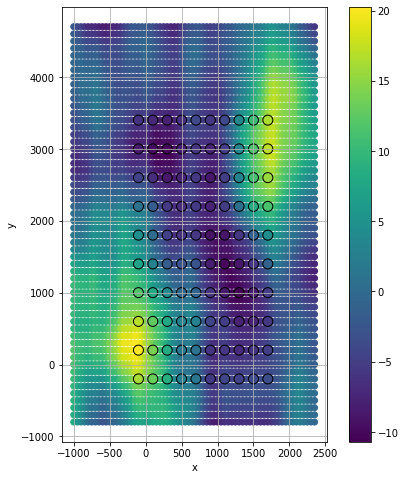
Condition and make predictions#
[11]:
plt.figure(figsize=(8, 8))
plt.ylabel('y')
plt.xlabel('x')
# xo = np.arange(0,dx*N,dx/3)[:,None]
xdo = np.arange(-dx*0.5*N, dx*1.2*N, dx/4)[:,None]
ydo = np.arange(-dy*0.2*N, dy*1.2*N, dy/4)[:,None]
# Make a grid
Xgo, Ygo = np.meshgrid(xdo, ydo)
# Vectorise grid and stack
Xvo = Xgo.ravel()
Yvo = Ygo.ravel()
Xo = np.hstack([Xvo[:,None], Yvo[:,None]])
OI = GPtideScipy(X, Xo, 0, covfunc, MAP[1:],
P=1, mean_func=None)
out_map = OI.conditional(zd)
plt.scatter(Xgo, Ygo, c=out_map, alpha=1)
plt.scatter(Xg, Yg, c=zg, s=100, edgecolors='k')
plt.grid()
plt.gca().set_aspect('equal')
plt.colorbar()
[11]:
<matplotlib.colorbar.Colorbar at 0x1ef276883a0>
[ ]: